- HOME
- Research
- Research Category
- Theoretical Chemistry Ishikita Laboratory
Theoretical Chemistry Ishikita Laboratory
Exploring mechanisms of proteins based on theoretical molecular chemistry to present a new strategy for molecular design and bioengineering
Understanding of the principles of protein function on the basis of the molecular structure
Proteins consist of only 20 types of amino acids, while they show large variety in their functions, e.g., redox activity, transporter, sensor, and antibodies. To clarify a relationship between functions and structures of proteins, we analyze molecular structures of proteins at the atomic level and calculate physical or chemical constants on the basis of theoretical chemistry.
Certainly, functions of proteins should be fully explained solely by the molecular structure even if the functions are seemingly complicated. “Just computing molecules” is not in our interest. Our mission is to uncover new but simple principles essential to the protein science through careful analysis of the target proteins. For example, we are trying to clarify the reaction mechanisms of natural photosynthetic proteins, e.g., O2- evolution, electron transfer, and proton transfer reactions. We also develop new tools for analysis of protein function.
Our challenges include:
(1) Toward understanding of functional mechanisms of proteins and macromolecules for molecular design
- ・Electron, proton, and energy transfer reactions in photosynthesis
- ・Correlation between structure and functions of photoreceptor and ion transporter
- ・Toward more active catalytic centers: elucidation of minimum key components that contribute to enzymatic reactions in enzymes
(2) Development of new chemical theories and computational methods
- ・Quantum mechanics model for molecular dynamic simulation
- ・Theoretical prediction of acid dissociation constants (pKa) by quantum chemical calculation
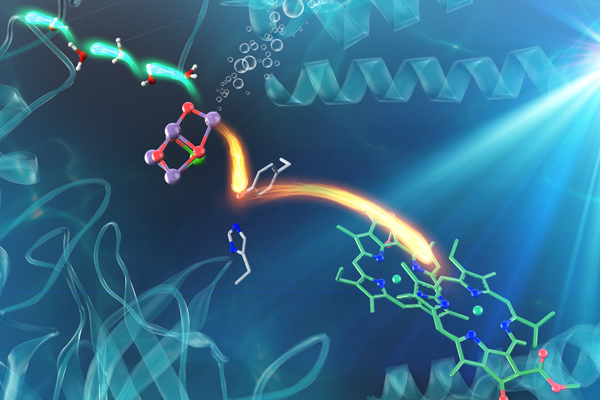
Electron and proton transfers in the water-oxidizing enzyme photosystem II

Super computer in our laboratory.
Exciting discussion.
My first encounter with photosynthesis research occurred when I went to Berlin to pursue my Ph.D. During my undergraduate and master's studies, I was interested in biomolecular devices. For example, the brain of a computer (the CPU) generates a lot of heat due to leakage current. With this background, I wanted to create biomolecular devices that surpassed silicon semiconductors, which led me to pursue an engineering degree. After being awarded a scholarship from Germany (DAAD), I was required to spend four months in Bremen for German language studies, which delayed the start of my research in Berlin. By the time I arrived in Berlin, the project I had described in my scholarship application was almost completed, and instead, I was given a new topic on electron transfer in photosynthetic reaction center proteins. I was instantly captivated by the crystal structure of the protein. The beauty of the symmetry in the arrangement of the pair of electron transfer pathways and the imbalance in the asymmetry of electron transfer activity. Solving this mystery—explaining it in terms of molecular chemistry that everyone can understand—has become my life's work.
Member

-
- Professor
Hiroshi ISHIKITA
Research Area: Biophysics - Professor

-
- Associate Professor
Keisuke SAITO
Research Area: Bio-and chemical physics - Associate Professor
Laboratory Homepage
Tags

